uva 1368 DNA Consensus String(检索)
来源:互联网 发布:剑三喵姐冷艳捏脸数据 编辑:程序博客网 时间:2024/04/29 21:57
Description
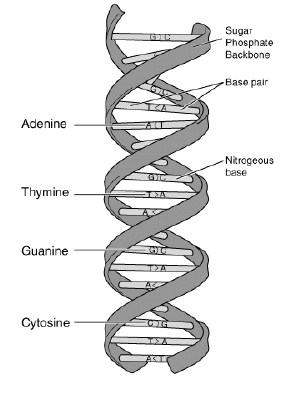 Figure 1.
Figure 1.
``Thymine-Adenine-Adenine-Cytosine-Thymine-Guanine-Cytosine-Cytosine-Guanine-Adenine-Thymine"
Then we can represent the above DNA strand with the string ``TAACTGCCGAT." The biologist Prof. Ahn found that a gene X commonly exists in the DNA strands of five different kinds of animals, namely dogs, cats, horses, cows, and monkeys. He also discovered that the DNA sequences of the gene X from each animal were very alike. See Figure 2.
Prof. Ahn thought that humans might also have the gene X and decided to search for the DNA sequence of X in human DNA. However, before searching, he should define a representative DNA sequence of gene X because its sequences are not exactly the same in the DNA of the five animals. He decided to use the Hamming distance to define the representative sequence. The Hamming distance is the number of different characters at each position from two strings of equal length. For example, assume we are given the two strings ``AGCAT" and ``GGAAT." The Hamming distance of these two strings is 2 because the 1st and the 3rd characters of the two strings are different. Using the Hamming distance, we can define a representative string for a set of multiple strings of equal length. Given a set of strings S = s1,..., sm of length n , the consensus error between a string y of length n and the set S is the sum of the Hamming distances between y and each si in S . If the consensus error between y and S is the minimum among all possible strings y of length n , y is called a consensus string of S . For example, given the three strings ``AGCAT" ``AGACT" and ``GGAAT" the consensus string of the given strings is ``AGAAT" because the sum of the Hamming distances between ``AGAAT" and the three strings is 3 which is minimal. (In this case, the consensus string is unique, but in general, there can be more than one consensus string.) We use the consensus string as a representative of the DNA sequence. For the example of Figure 2 above, a consensus string of gene X is ``GCAAATGGCTGTGCA" and the consensus error is 7.
Input
Output
Sample Input
3 5 8 TATGATAC TAAGCTAC AAAGATCC TGAGATAC TAAGATGT 4 10 ACGTACGTAC CCGTACGTAG GCGTACGTAT TCGTACGTAA 6 10 ATGTTACCAT AAGTTACGAT AACAAAGCAA AAGTTACCTT AAGTTACCAA TACTTACCAA
Sample Output
TAAGATAC 7 ACGTACGTAA 6 AAGTTACCAA 12
题目大意:找出一个串,要求和所有给出的字符串汉明距离最小(字典序也要求最小)
解题思路:遍历每个字符,将所有字符串中同一位的字符统计一遍,记录最多的字符赋值在所求串上。
#include<stdio.h>#include<string.h>#define N 1005#define M 55char str[M][N];char tem[N];int num[M];int m, n;char find(int k){int t[30];memset(t, 0, sizeof(t));for (int i = 0; i < m; i++)t[str[i][k] - 'A']++;int cnt = 0, x = 0;for (int i = 0; i < 26; i++)if (cnt < t[i]){x = i;cnt = t[i];}return 'A' + x;}int main(){int t;scanf("%d", &t);while (t--){// Init.memset(str, 0, sizeof(str));memset(tem, 0, sizeof(tem));memset(num, 0, sizeof(num));// Read.scanf("%d%d%*c", &m, &n);for (int i = 0; i < m; i++)gets(str[i]);// Handle.int cnt = 0;for (int i = 0; i < n; i++){char c = find(i);tem[i] = c;for (int j = 0; j < m; j++)if (str[j][i] != c)cnt++;}tem[n] = '\0';puts(tem);printf("%d\n", cnt);}return 0;}- uva 1368 DNA Consensus String(检索)
- uva 1368 - DNA Consensus String(贪心)
- uva - 1368 - DNA Consensus String(字符串)
- UVa 1368 - DNA Consensus String(贪心)
- uva 1368 - DNA Consensus String(贪心)
- (入门)uva 1368 DNA Consensus String
- UVA 1368 - DNA Consensus String
- uva 1368 DNA Consensus String
- Uva-1368-DNA Consensus String
- UVa 1368 DNA Consensus String
- UVA 1368 DNA Consensus String
- UVA 1368 DNA Consensus String
- UVa:1368 DNA Consensus String
- uva 1368 - DNA Consensus String
- UVa 1368 - DNA Consensus String
- UVa 1368 - DNA Consensus String
- UVa 1368 DNA Consensus String
- UVa 1368 - DNA Consensus String
- OK6410启动代码(3)
- 开始C/C++生涯
- OK6410启动代码(4)
- iOS extracts: Adding Color
- windows 安装 flash builder 开发flex 4 项目
- uva 1368 DNA Consensus String(检索)
- Html5问题备忘
- 8. 童子军规则
- 程序猿的第一步
- .net中将DataGridView内的数据导出为Excel表格
- HDU4630 No Pain No Game
- C# 中的委托和事件
- 黑马程序员---银行业务调度系统
- 底部能带消息提示的TabHost的实


