Seoul 2006 / UVa 1368 DNA Consensus String (字符串处理)
来源:互联网 发布:工资查询软件 编辑:程序博客网 时间:2024/05/21 18:48
1368 - DNA Consensus String
Time limit: 3.000 seconds
http://uva.onlinejudge.org/index.php?option=com_onlinejudge&Itemid=8&category=457&page=show_problem&problem=4114
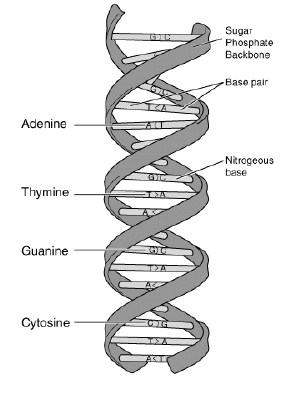 Figure 1.
Figure 1.
``Thymine-Adenine-Adenine-Cytosine-Thymine-Guanine-Cytosine-Cytosine-Guanine-Adenine-Thymine"
Then we can represent the above DNA strand with the string ``TAACTGCCGAT." The biologist Prof. Ahn found that a gene X commonly exists in the DNA strands of five different kinds of animals, namely dogs, cats, horses, cows, and monkeys. He also discovered that the DNA sequences of the gene X from each animal were very alike. See Figure 2.
Prof. Ahn thought that humans might also have the gene X and decided to search for the DNA sequence of X in human DNA. However, before searching, he should define a representative DNA sequence of gene X because its sequences are not exactly the same in the DNA of the five animals. He decided to use the Hamming distance to define the representative sequence. The Hamming distance is the number of different characters at each position from two strings of equal length. For example, assume we are given the two strings ``AGCAT" and ``GGAAT." The Hamming distance of these two strings is 2 because the 1st and the 3rd characters of the two strings are different. Using the Hamming distance, we can define a representative string for a set of multiple strings of equal length. Given a set of strings S = s1,..., sm of length n , the consensus error between a string y of length n and the set S is the sum of the Hamming distances between y and each si in S. If the consensus error between y and S is the minimum among all possible strings y of length n , y is called a consensus string of S . For example, given the three strings ``AGCAT" ``AGACT" and ``GGAAT" the consensus string of the given strings is ``AGAAT" because the sum of the Hamming distances between ``AGAAT" and the three strings is 3 which is minimal. (In this case, the consensus string is unique, but in general, there can be more than one consensus string.) We use the consensus string as a representative of the DNA sequence. For the example of Figure 2 above, a consensus string of gene X is ``GCAAATGGCTGTGCA" and the consensus error is 7.
Input
Your program is to read from standard input. The input consists of T test cases. The number of test cases Tis given in the first line of the input. Each test case starts with a line containing two integers m and nwhich are separated by a single space. The integer m (4Output
Your program is to write to standard output. Print the consensus string in the first line of each case and the consensus error in the second line of each case. If there exists more than one consensus string, print the lexicographically smallest consensus string. The following shows sample input and output for three test cases.Sample Input
3 5 8 TATGATAC TAAGCTAC AAAGATCC TGAGATAC TAAGATGT 4 10 ACGTACGTAC CCGTACGTAG GCGTACGTAT TCGTACGTAA 6 10 ATGTTACCAT AAGTTACGAT AACAAAGCAA AAGTTACCTT AAGTTACCAA TACTTACCAA
Sample Output
TAAGATAC 7 ACGTACGTAA 6 AAGTTACCAA 12
water.
完整代码:
/*0.012s*/#include<cstdio>const char ACGT[4] = {'A', 'C', 'G', 'T'};///已经保证了字典序char str[50][1001];struct DNA{char ch;int c;} cnt[4], maxn;int main(void){for (int i = 0; i < 4; ++i)cnt[i].ch = ACGT[i];int t;scanf("%d", &t);while (t--){int m, n, error = 0;scanf("%d%d", &m, &n);getchar();for (int i = 0; i < m; ++i)gets(str[i]);for (int j = 0; j < n; ++j){for (int i = 0; i < 4; ++i)cnt[i].c = 0;for (int i = 0; i < m; ++i){if (str[i][j] == 'A') ++cnt[0].c;else if (str[i][j] == 'C') ++cnt[1].c;else if (str[i][j] == 'G') ++cnt[2].c;else ++cnt[3].c;}maxn = cnt[0];for (int i = 1; i < 4; ++i)if (cnt[i].c > maxn.c)maxn = cnt[i];putchar(maxn.ch);error += m - maxn.c;}printf("\n%d\n", error);}return 0;}- Seoul 2006 / UVa 1368 DNA Consensus String (字符串处理)
- UVA 1368 DNA Consensus String【ACM/ICPC Seoul 2006】
- DNA序列(DNA Consensus String, ACM/ICPC seoul 2006, UVa 1368)
- uva 1368 - DNA Consensus String(字符串处理)
- UVA - 1368 - DNA Consensus String (字符串处理)
- UVA - 1368 DNA Consensus String :简单字符串处理
- uva - 1368 - DNA Consensus String(字符串)
- UVa 1368 - DNA Consensus String【字符串】
- UVa 1368 DNA Consensus String 【字符串】【贪心】
- DNA序列 (DNA Consensus String, ACM/ICPC Seoul 2006 UVa1368)
- UVA 1368 - DNA Consensus String
- uva 1368 DNA Consensus String
- Uva-1368-DNA Consensus String
- UVa 1368 DNA Consensus String
- UVA 1368 DNA Consensus String
- UVA 1368 DNA Consensus String
- UVa:1368 DNA Consensus String
- uva 1368 - DNA Consensus String
- ViewPager 滑动切换 activity
- cocos2d-x win32移植到android
- Windows下编辑的txt在linux下乱码的解决办法
- 用封装的方法实现从文件夹名下所有的指定类型文件数据导入到数据库
- box2d弹球 cocos2d-x重力感应(cocos2d-x2.1)
- Seoul 2006 / UVa 1368 DNA Consensus String (字符串处理)
- iOS 实现集成话分享
- 博客搬家
- myecplice2013 破解 没有common文件夹
- ViewPager动态加载数据
- thrift shows CLOSE_WAIL error
- Nginx如何处理一个请求
- ExtJS4.2学习(6)——基础知识之proxy篇
- Android 杀不掉的后台服务的一种实现


